-Search query
-Search result
Showing 1 - 50 of 4,717 items for (author: ali & s)

EMDB-44372: 
In-cell Saccharomyces cerevisiae nuclear pore complex with single nuclear ring
Method: subtomogram averaging / : Singh D, Hutchings J, Villa E

EMDB-44377: 
In-cell Saccharomyces cerevisiae nuclear pore complex with double nuclear ring and basket
Method: subtomogram averaging / : Singh D, Hutchings J, Villa E

EMDB-44379: 
In-cell Mus musculus nuclear pore complex with nuclear basket
Method: subtomogram averaging / : Hutchings J, Singh D, Villa E

EMDB-44381: 
In-cell Toxoplasma gondii nuclear pore complex
Method: subtomogram averaging / : Singh D, Hutchings J, Li Z, Guo Q, Villa E

EMDB-45197: 
In-cell Saccharomyces cerevisiae symmetry-expanded nuclear pore complex with double nuclear ring and basket
Method: subtomogram averaging / : Singh D, Hutchings J, Villa E

EMDB-45198: 
In-cell Saccharomyces cerevisiae symmetry-expanded nuclear pore complex with single nuclear ring
Method: subtomogram averaging / : Singh D, Hutchings J, Villa E

EMDB-45199: 
In-cell Saccharomyces cerevisiae nuclear pore complex cytoplasmic ring focused refinement
Method: subtomogram averaging / : Singh D, Hutchings J, Villa E

EMDB-45200: 
In-cell Saccharomyces cerevisiae nuclear pore complex inner ring focused refinement
Method: subtomogram averaging / : Singh D, Hutchings J, Villa E

EMDB-45201: 
In-cell Saccharomyces cerevisiae nuclear pore complex single nuclear ring focused refinement
Method: subtomogram averaging / : Singh D, Hutchings J, Villa E

EMDB-45202: 
In-cell Saccharomyces cerevisiae nuclear pore complex double nuclear ring focused refinement
Method: subtomogram averaging / : Singh D, Hutchings J, Villa E

EMDB-45203: 
In-cell Saccharomyces cerevisiae nuclear pore complex nuclear basket focused refinement
Method: subtomogram averaging / : Singh D, Hutchings J, Villa E

EMDB-45204: 
In-cell Saccharomyces cerevisiae nuclear pore complex membrane focused refinement for single nuclear ring
Method: subtomogram averaging / : Singh D, Hutchings J, Villa E

EMDB-45205: 
In-cell Saccharomyces cerevisiae nuclear pore complex membrane focused refinement for double nuclear ring
Method: subtomogram averaging / : Singh D, Hutchings J, Villa E

EMDB-45216: 
In-cell Mus musculus nuclear pore complex with nuclear basket consensus map
Method: subtomogram averaging / : Hutchings J, Singh D, Villa E

EMDB-45219: 
In-cell Mus musculus nuclear pore complex cytoplasmic ring focused refinement
Method: subtomogram averaging / : Hutchings J, Singh D, Villa E

EMDB-45220: 
In-cell Mus musculus nuclear pore complex inner ring focused refinement
Method: subtomogram averaging / : Hutchings J, Singh D, Villa E

EMDB-45222: 
In-cell Mus musculus nuclear pore complex nuclear ring focused refinement
Method: subtomogram averaging / : Hutchings J, Singh D, Villa E

EMDB-45223: 
In-cell Mus musculus nuclear pore complex basket focused refinement
Method: subtomogram averaging / : Hutchings J, Singh D, Villa E

EMDB-45227: 
In-cell Mus musculus nuclear pore complex membrane focused refinement
Method: subtomogram averaging / : Hutchings J, Singh D, Villa E

EMDB-45228: 
In-cell Toxoplasma gondii symmetry-expanded nuclear pore complex
Method: subtomogram averaging / : Singh D, Hutchings J, Li Z, Guo Q, Villa E
EMDB-45255: 
In-cell Saccharomyces cerevisiae C8-symmetrised nuclear pore complex consensus map
Method: subtomogram averaging / : Singh D, Hutchings J, Villa E

EMDB-45256: 
In-cell Saccharomyces cerevisiae symmetry-expanded nuclear pore complex consensus map
Method: subtomogram averaging / : Singh D, Hutchings J, Villa E

EMDB-45257: 
In-cell Mus musculus nuclear pore complex with nuclear basket consensus map
Method: subtomogram averaging / : Hutchings J, Singh D, Villa E

EMDB-45258: 
In-cell Mus musculus symmetry-expanded nuclear pore complex with nuclear basket consensus map
Method: subtomogram averaging / : Hutchings J, Singh D, Villa E

EMDB-45259: 
In-cell Toxoplasma gondii C8-symmetrised nuclear pore complex consensus map
Method: subtomogram averaging / : Singh D, Hutchings J, Li Z, Guo Q, Villa E

EMDB-42291: 
Structure of the human INTS9-INTS11-BRAT1 complex
Method: single particle / : Lin M, Tong L

EMDB-42292: 
Structure of the Drosophila IntS11-CG7044(dBRAT1) complex
Method: single particle / : Lin M, Tong L
EMDB-18621: 
ASCT2 trimer in lipid nanodiscs with bound glutamine and Na+ ions in the outward-facing state (OFS)
Method: single particle / : Borowska A, Rheinberger J, Paulino C, Slotboom DJ
EMDB-18622: 
ASCT2 protomer in lipid nanodiscs with bound glutamine and Na+ ions in the outward-facing state (OFS.1)
Method: single particle / : Borowska A, Rheinberger J, Paulino C, Slotboom DJ
EMDB-18623: 
ASCT2 protomer in lipid nanodiscs with bound glutamine and Na+ ions in the outward-facing state (OFS.2)
Method: single particle / : Borowska A, Rheinberger J, Paulino C, Slotboom DJ
EMDB-18624: 
ASCT2 protomer in lipid nanodiscs with bound glutamine and Na+ ions in the outward-facing state (OFS.3)
Method: single particle / : Borowska A, Rheinberger J, Paulino C, Slotboom DJ
EMDB-18625: 
ASCT2 protomer in lipid nanodiscs with bound glutamine and Na+ ions in the intermediate outward-facing state (iOFS-up)
Method: single particle / : Borowska A, Rheinberger J, Paulino C, Slotboom DJ
EMDB-18626: 
ASCT2 protomer in lipid nanodiscs with bound glutamine and Na+ ions in the intermediate outward-facing state (iOFS-down)
Method: single particle / : Borowska A, Rheinberger J, Paulino C, Slotboom DJ
EMDB-18627: 
ASCT2 protomer in lipid nanodiscs under low Na+ concentration in the outward-facing state (OFS)
Method: single particle / : Borowska A, Rheinberger J, Paulino C, Slotboom DJ
EMDB-18628: 
ASCT2 protomer in lipid nanodiscs under low Na+ concentration in the intermediate outward-facing state (iOFS-up)
Method: single particle / : Borowska A, Rheinberger J, Paulino C, Slotboom DJ

PDB-8qro: 
ASCT2 trimer in lipid nanodiscs with bound glutamine and Na+ ions in the outward-facing state (OFS)
Method: single particle / : Borowska A, Rheinberger J, Paulino C, Slotboom DJ

PDB-8qrp: 
ASCT2 protomer in lipid nanodiscs with bound glutamine and Na+ ions in the outward-facing state (OFS.1)
Method: single particle / : Borowska A, Rheinberger J, Paulino C, Slotboom DJ

PDB-8qrq: 
ASCT2 protomer in lipid nanodiscs with bound glutamine and Na+ ions in the outward-facing state (OFS.2)
Method: single particle / : Borowska A, Rheinberger J, Paulino C, Slotboom DJ

PDB-8qrr: 
ASCT2 protomer in lipid nanodiscs with bound glutamine and Na+ ions in the outward-facing state (OFS.3)
Method: single particle / : Borowska A, Rheinberger J, Paulino C, Slotboom DJ

PDB-8qrs: 
ASCT2 protomer in lipid nanodiscs with bound glutamine and Na+ ions in the intermediate outward-facing state (iOFS-up)
Method: single particle / : Borowska A, Rheinberger J, Paulino C, Slotboom DJ

PDB-8qru: 
ASCT2 protomer in lipid nanodiscs with bound glutamine and Na+ ions in the intermediate outward-facing state (iOFS-down)
Method: single particle / : Borowska A, Rheinberger J, Paulino C, Slotboom DJ

PDB-8qrv: 
ASCT2 protomer in lipid nanodiscs under low Na+ concentration in the outward-facing state (OFS)
Method: single particle / : Borowska A, Rheinberger J, Paulino C, Slotboom DJ

PDB-8qrw: 
ASCT2 protomer in lipid nanodiscs under low Na+ concentration in the intermediate outward-facing state (iOFS-up)
Method: single particle / : Borowska A, Rheinberger J, Paulino C, Slotboom DJ
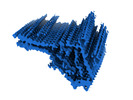

EMDB-50272: 
Additional cryo-EM structure of cardiac amyloid AL59 - mixed polymorph
Method: helical / : Schulte T, Speranzini V, Chaves-Sanjuan A, Milazzo M, Ricagno S

EMDB-50271: 
Additional cryo-EM structure of cardiac amyloid AL59 - bent polymorph
Method: helical / : Schulte T, Speranzini V, Chaves-Sanjuan A, Milazzo M, Ricagno S

EMDB-51031: 
Cryo-electron tomogram of AL59 amyloids interacting with collagen VI
Method: electron tomography / : Sicking K, Fernandez-Busnadiego R, Ricagno S

EMDB-51032: 
Cryo-electron tomogram of AL59 amyloids interacting with collagen VI
Method: electron tomography / : Sicking K, Fernandez-Busnadiego R, Ricagno S

EMDB-51033: 
Cryo-electron tomogram of AL59 amyloids interacting with collagen VI
Method: electron tomography / : Sicking K, Fernandez-Busnadiego R, Ricagno S

EMDB-51038: 
Cryo-electron tomogram of AL59 amyloids interacting with collagen VI
Method: electron tomography / : Sicking K, Fernandez-Busnadiego R, Ricagno S

EMDB-19397: 
Composite map of the C. elegans Intron Lariat Spliceosome primed for disassembly (ILS')
Method: single particle / : Vorlaender MK, Rothe P, Plaschka C
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
